Computer Facilities: Brain Imaging Core computing facilities consist of centralized computing infrastructure, a shared central compute resource, a walk-in image analysis bay, and desktop/laptop computers for BI lab personnel.
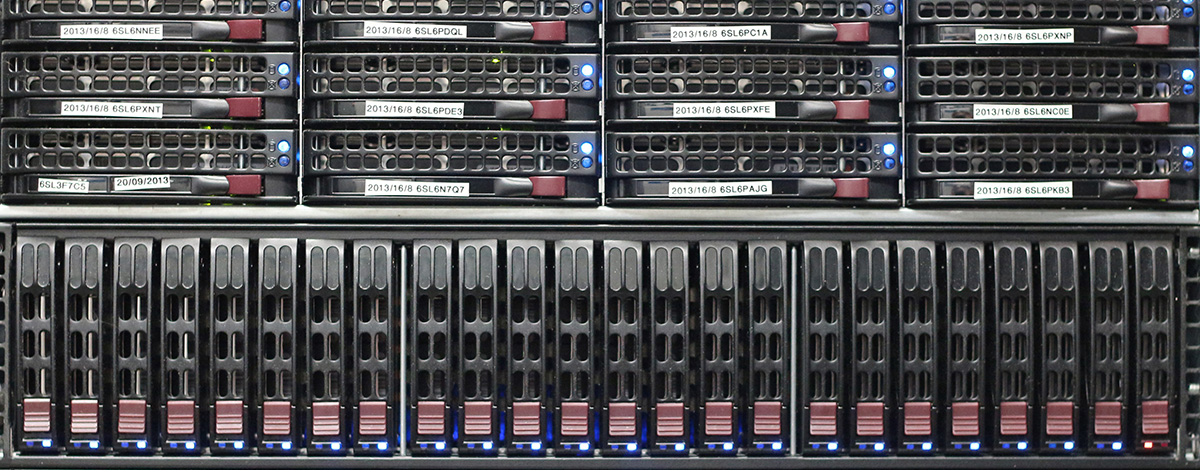
The centralized computing infrastructure is housed in a dedicated computer room with raised floor, air conditioning, and APC uninterruptable power supplies. Central storage consists of 111TB of network attached, 10k/15k rpm Serial Attached SCSI RAID6 disk arrays. Ancillary file servers provide additional network attached storage in a variety of RAID0/1/6, configurations. Total network storage is 142TB. Data is made available to all Core workstations via nfs and smb/cifs protocols. Backups are controlled by EMC Legato enterprise backup software to an HP StorageWorks MSL8096 library with three LTO5 tape drives on a SAN fabric in a drive sharing configuration. Total tape capacity is approximately two PetaBytes. The windows active directory domain is served by 3 quad-code Xeon servers with fully redundant hardware capabilities. A web application platform provides internet-based research paradigm presentation. Other servers provide web, database, monitoring, alerting, windows remote desktop, and other core network services.
A shared central computing resource is housed in the dedicated computer room. This resource provides 288 computing cores, approximately 2 TB of aggregate RAM, local scratch disk, is accessible to lab users both locally and over the internet, and has full connectivity to all Brain Imaging Core computing resources. The Condor high-throughput computing environment is used to manage resource allocation and scheduling. NVIDIA Tesla K80 and Titan Xp graphical processors provide GPU acceleration for CUDA-enabled processing pipelines. A separate, dedicated condor submit node provides access to 7000+ CPUs via the UW-Madison Center for High Throughput computing.
The walk-in image analysis bay provides 12 shared OS X and windows workstations for Brain Imaging lab users, along with monitor/keyboard/mouse stations for laptop users.
The Brain Imaging Core also supports approximately 250 desktop, laptop, instrumentation, and data collection computers, running Scientific Linux, OS X/macOS, and windows operating systems. Computers within the facility have full connectivity to the Core storage, compute, and software resources.
Access to a wide variety of data analysis software is available throughout the Brain Imaging Core. Most software packages are available on every computer in the Core. The following programming languages are actively used and supported: C/C++, Java, Perl, Python, Ruby, TCL, IDL (Research Systems, Inc.), and Matlab (MathWorks). The Core uses standard software for writing to CD/RW media, as well as a customized in-house program for performing robust data backup and archival to CD/RW disks. Statistical analysis packages include SPSS, R, Matlab, SAS, and Mathematica. Machine shop drawings use AutoCAD. The Core includes several methodological research groups who vigorously produce in-house software to aid their investigations related to MRI, PET, and EEG, although where possible, the Core uses existing software obtained either commercially or from other academic research groups.
MRI: Stimuli are presented by one of several software packages including E-Prime, PsychoPy, Presentation, and PsychToolbox. Many software packages are available for processing and analysis of MRI data, including FSL, AFNI, SPM, FreeSurfer, ANTs, fmriprep, camino, and DTI-TK, with other packages installed by researcher requests. Advanced analysis and display are performed by in-house developed tools.
PET: Manufacturers software is used for acquisition and reconstruction of PET data from the ECAT V7.2.2 and the Concorde/microPET; in-house software (Spamalize) is used for reconstruction of data from an offsite PET scanner. Images are coregistered and realigned using publicly available software (e.g SPM, FLIRT) or an in-house program (BrainSqueezer). Kinetic analysis for the measurement of quantitative metrices is performed on both ROI- and voxel-level data using in-house software developed in the MATLAB environment. Quantitative analysis is frequently performed for evaluation of cerebral perfusion and metabolism, oxygen metabolism, neuroreceptor binding and assessment of neuroinflammation, amyloid and tau protein burden.
Psychophysiology: PST E-Prime software controls stimulus presentation. BIOPAC AcqKnowledge and Tobii Studio software are used for data acquisition. Data preprocessing and analysis are performed using sophisticated in-house statistical tools developed in MATLAB, Python, and R.
Image Display: A variety of image display programs are available, including AFNI, SPM, FSL, Freesurfer, Spamalize, and, CURRY.